# Quantile function for a Weibull distribution with a cure proportion
qWeibullCure <- function(p, p0, shape = 1, scale){
res <- rep(NA, length(p))
ind1 <- (p <= p0)
ind2 <- (p > p0)
res[ind1] <- Inf
res[ind2] <- qweibull(1 - (p[ind2] - p0) / (1 - p0), shape = shape, scale = scale)
return(res)
}
# censored time-to-event data with censoring after pre-specified number of events
rWeibull1arm <- function(shape = 1, scale, cure = 0, recruit, dropout = 0, start.accrual = 0, cutoff, seed = NA){
# shape Weibull shape parameter.
# scale: Weibull scale parameter
# cure: Proportion of patients assumed to be cured, i.e. with an event at +infty.
# recruit: Recruitment.
# dropout: Drop-out rate, on same time scale as med.
# start.accrual: Time unit where accrual should start. Might be useful when simulating multi-stage trials.
# cutoff: Cutoff, #events the final censored data should have (can be a vector of multiple cutoffs).
# seed: If different from NA, seed used to generate random numbers.
#
# Kaspar Rufibach, June 2014
if (is.na(seed) == FALSE){set.seed(seed)}
n <- sum(recruit)
# generate arrival times
arrive <- rep(1:length(recruit), times = recruit)
arrivetime <- NULL
for (i in 1:n){arrivetime[i] <- runif(1, min = arrive[i] - 1, max = arrive[i])}
arrivetime <- start.accrual + sort(arrivetime)
# generate event times: Exp(lambda) = Weibull(shape = 1, scale = 1 / lambda)
eventtime <- qWeibullCure(runif(n), p0 = cure, shape = shape, scale = scale)
# Apply drop-out. Do this before applying the cutoff below, in order to correctly count necessary #events.
dropouttime <- rep(Inf, n)
if (dropout > 0){dropouttime <- rexp(n, rate = dropout)}
event.dropout <- ifelse(eventtime > dropouttime, 0, 1)
time.dropout <- ifelse(event.dropout == 1, eventtime, dropouttime)
# observed times, taking into account staggered entry
tottime <- arrivetime + eventtime
# find cutoff based on number of targeted events
# only look among patients that are not considered dropped-out
time <- data.frame(matrix(NA, ncol = length(cutoff), nrow = n))
event <- time
cutoff.time <- rep(NA, length(cutoff))
for (j in 1:length(cutoff)){
cutoff.time[j] <- sort(tottime[event.dropout == 1])[cutoff[j]]
# apply administrative censoring at cutoff
event[event.dropout == 1, j] <- ifelse(tottime[event.dropout == 1] > cutoff.time[j], 0, 1)
event[event.dropout == 0, j] <- 0
# define time to event, taking into account both types of censoring
time[event.dropout == 1, j] <- ifelse(event[, j] == 1, eventtime, cutoff.time[j] - arrivetime)[event.dropout == 1] # same as: pmin(tottime, cutoff.time) - arrive
time[event.dropout == 0, j] <- pmin(cutoff.time[j] - arrivetime, time.dropout)[event.dropout == 0]
# remove times for patients arriving after the cutoff
rem <- (arrivetime > cutoff.time[j])
if (TRUE %in% rem){time[rem, j] <- NA}
}
# generate output
tab <- data.frame(cbind(1:n, arrivetime, eventtime, tottime, dropouttime, time, event))
colnames(tab) <- c("pat", "arrivetime", "eventtime", "tottime", "dropouttime", paste("time cutoff = ", cutoff, sep = ""), paste("event cutoff = ", cutoff, sep = ""))
res <- list("cutoff.time" = cutoff.time, "tab" = tab)
return(res)
}
# censored time-to-event data with censoring after pre-specified number of events, for two treatment arms
rWeibull2arm <- function(shape = c(1, 1), scale, cure = c(0, 0), recruit, dropout = c(0, 0), start.accrual = c(0, 0), cutoff, seed = NA){
# shape 2-d vector of Weibull shape parameter.
# scale 2-d vector of Weibull scale parameter.
# cure: 2-d vector with cure proportion assumed in each arm.
# recruit: List with two elements, vector of recruitment in each arm.
# dropout: 2-d vector with drop-out rate for each arm, on same time scale as med.
# start.accrual: 2-d vector of time when accrual should start. Might be useful when simulating multi-stage trials.
# cutoff: Cutoff, #events the final censored data should have (can be a vector of multiple cutoffs).
# seed: If different from NA, seed used to generate random numbers.
#
# Kaspar Rufibach, June 2014
if (is.na(seed) == FALSE){set.seed(seed)}
dat1 <- rWeibull1arm(scale = scale[1], shape = shape[1], recruit = recruit[[1]], cutoff = 1,
dropout = dropout[1], cure = cure[1], start.accrual = start.accrual[1], seed = NA)$tab
dat2 <- rWeibull1arm(scale = scale[2], shape = shape[2], recruit = recruit[[2]], cutoff = 1,
dropout = dropout[2], cure = cure[2], start.accrual = start.accrual[2], seed = NA)$tab
n <- c(nrow(dat1), nrow(dat2))
# treatment variable
tmt <- factor(c(rep(0, n[1]), rep(1, n[2])), levels = 0:1, labels = c("A", "B"))
arrivetime <- c(dat1[, "arrivetime"], dat2[, "arrivetime"])
eventtime <- c(dat1[, "eventtime"], dat2[, "eventtime"])
tottime <- c(dat1[, "tottime"], dat2[, "tottime"])
dropouttime <- c(dat1[, "dropouttime"], dat2[, "dropouttime"])
# Apply drop-out. Do this before applying the cutoff below, in order to correctly count necessary #events.
event.dropout <- ifelse(eventtime > dropouttime, 0, 1)
time.dropout <- ifelse(event.dropout == 1, eventtime, dropouttime)
# find cutoff based on number of targeted events
# only look among patients that are not considered dropped-out
time <- data.frame(matrix(NA, ncol = length(cutoff), nrow = sum(n)))
event <- time
cutoff.time <- rep(NA, length(cutoff))
for (j in 1:length(cutoff)){
cutoff.time[j] <- sort(tottime[event.dropout == 1])[cutoff[j]]
# apply administrative censoring at cutoff
event[event.dropout == 1, j] <- ifelse(tottime[event.dropout == 1] > cutoff.time[j], 0, 1)
event[event.dropout == 0, j] <- 0
# define time to event, taking into account both types of censoring
time[event.dropout == 1, j] <- ifelse(event[, j] == 1, eventtime, cutoff.time[j] - arrivetime)[event.dropout == 1]
time[event.dropout == 0, j] <- pmin(cutoff.time[j] - arrivetime, time.dropout)[event.dropout == 0]
# remove times for patients arriving after the cutoff
rem <- (arrivetime > cutoff.time[j])
if (TRUE %in% rem){time[rem, j] <- NA}
}
# generate output
tab <- data.frame(cbind(1:sum(n), tmt, arrivetime, eventtime, tottime, dropouttime, time, event))
colnames(tab) <- c("pat", "tmt", "arrivetime", "eventtime", "tottime", "dropouttime",
paste("time cutoff = ", cutoff, sep = ""), paste("event cutoff = ", cutoff, sep = ""))
res <- list("cutoff.time" = cutoff.time, "tab" = tab)
return(res)
}
# functions to plot results
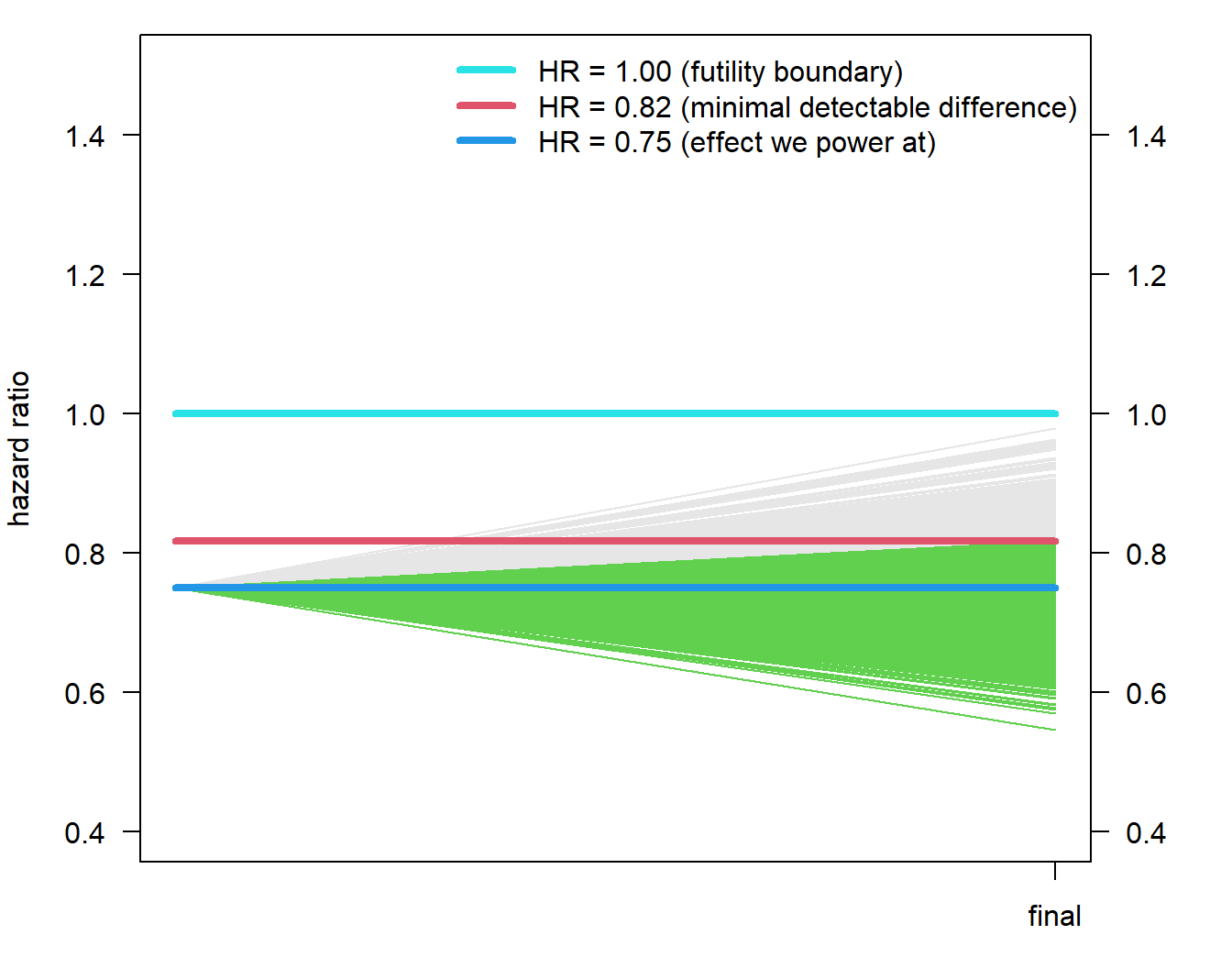
horiz <- function(vert = FALSE){
segments(0, 1, 1, 1, col = 5, lwd = 4, lty = 1)
segments(0, hr, 1, hr, col = 4, lwd = 4, lty = 1)
segments(0, mdd_no_interim, 1, mdd_no_interim, col = 2, lwd = 4, lty = 1)
legend("topright", paste("HR = ", disp(c(1, mdd_no_interim, hr), 2), " (",
c("futility boundary", "minimal detectable difference", "effect we power at"), ")",
sep = ""), bty = "n", lty = 1, col = c(5, 2, 4), lwd = 4)
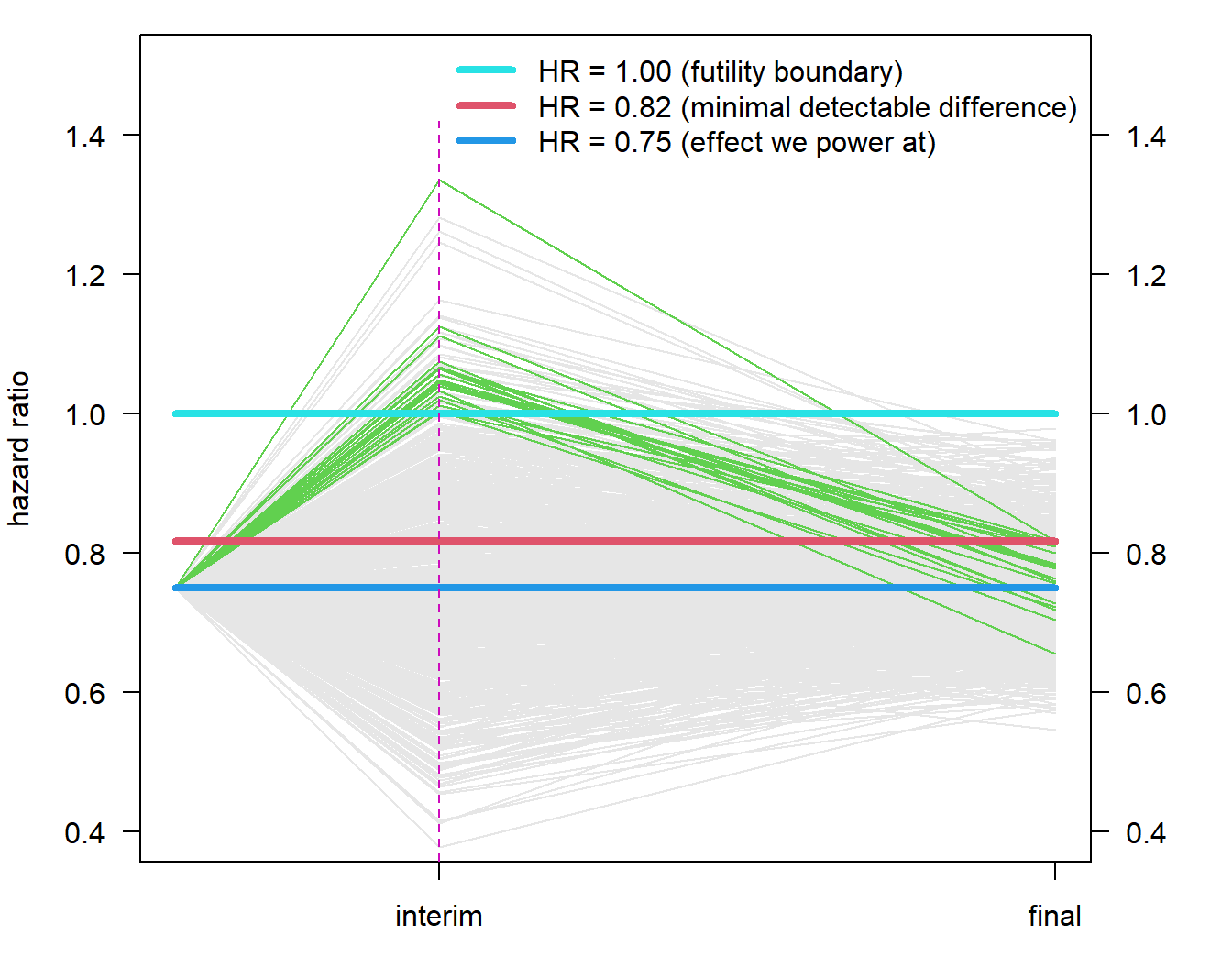
# vertical line
if (vert){segments(inter_x[1], 0, inter_x[1], 1.42, lty = 2, col = 6)}
}
plot_empty <- function(){
inter_x <- cutoff / max(cutoff)
yli_traj <- c(0.4, 1.5)
par(las = 1)
par(mar = c(3, 4, 1, 4), las = 1)
plot(0, 0, type = "n", xlim = c(0, 1), ylim = yli_traj, xlab = "", xaxt = "n", ylab = "hazard ratio")
axis(side = 4, at = seq(0, 2, by = 0.2), labels = disp(seq(0, 2, by = 0.2), 1))
axis(side = 1, at = inter_x[2], labels = c("interim", "final")[2])
}